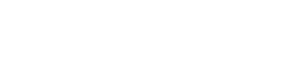Five Opportunities at the Interface Between Structural Biology and AI, and their Impact on Drug Discovery
by Denis Bucher, Principal Scientist at leadXpro and Head of Molecular Modelling and Design
Drug Discovery will be impacted by the advent of AI, but how?
To answer this question we have selected 5 fast-growing trends at the interface between structural biology and AI that promise to have high impact on drug discovery in the next few years.
1. Designing Ligands de novo
The most common origin of small molecule therapeutics has been compound-screening, done using experimental libraries. Today’s AI enables better in-silico selection of screening libraries and de novo design of compounds. De novo drug design is an iterative process in which the three-dimensional structure of the receptor is used to design new molecules from scratch (Fig. 1). Historically the design of molecules has been done by trained experts, but AI is now able to generate successful molecular ideas under a set of constraints.
An interesting approach consists in combining different neural nets (NNs) trained for specific tasks (Paul et al., 2021; Chan et al., 2019) in order to automate all the required steps of molecular design in silico before testing them experimentally. For instance:
- De novo design of molecules using knowledge gathered from previous rounds (Olivecrona et al., 2017; Schneider and Fechner, 2005)
- Prediction of synthetic feasibility (de Almeida et al., 2019; Struble et al., 2020)
- Prediction of ADME/tox (requires sufficient high-quality training data)
- Prediction of affinity using 3D protein-ligand structures => go back to (1)
Here, high quality 3D structures of the protein-ligand complexes are needed to enable the rapid in-silico design of potent compounds.
AI facilitates all the remaining steps, which leads to an accelerated identification of preclinical drug candidates.
2. Designing Proteins
Protein engineering – fusion protein insertion, deletions, and mutations – is a common approach in structural biology to enhance expression and purification yields and produce proteins that are conformationally more stable, and therefore, more amenable to high-resolution structural work by X-ray crystallography or Cryo-EM. For example, in a recent contribution by leadXpro (Botte et al., 2022) – nanobodies were specifically designed as chaperones to solve the structure of the bacterial LptDE transporter by CryoEM. (see leadXpro news article)
With the release of AI empowered software programs, such as Alphafold 2 (AF2) (Jumper et al., 2021), designing proteins for structural resolution has become more feasible and accurate. One key concept behind AF2 is the idea that residues in proteins located in proximity in 3-dimensional space have co-evolved. Using the exceptional ability of AI to learn meaningful patterns in multiple sequence alignments (MSAs), AF2 provides accurate predictions for apo (ligand-free) structures that rival the accuracy of experimental data. Furthermore, other software like Rosetta or MD simulations, provide complementary information about the protein model free energy (i.e., its stability).
However, an important current limitation of Alphafold 2 is the inability to predict protein-ligand complex structures, and conformational changes upon ligand binding, which still needs to be done experimentally.
3. Predicting Protein Stability and Expression
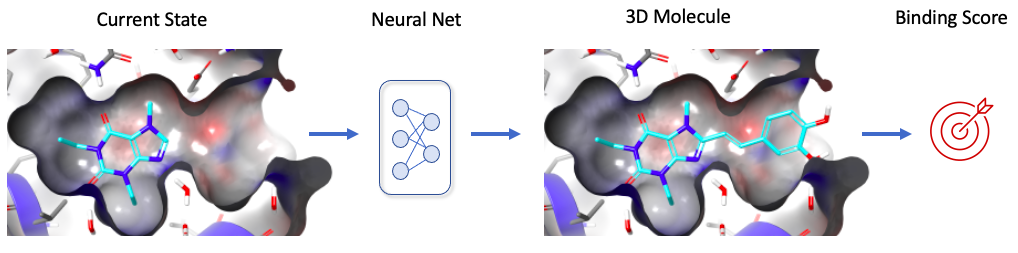
AI can combine data from different sources, detect complex patterns, and prioritize information in context. This is often better suited than conventional machine learning techniques (or human learning) to detect mutations that can significantly improve the stability of proteins, and therefore the probability of solving the structure at high-resolution. Advanced algorithms are being explored to combine in an optimal way information from literature and patent publications, relevant patterns found in MSA (2D sequence alignments) (Hopf et al., 2019), as well as the 3D environment of residues in protein models (Fig. 2) (Popov et al., 2018). For predicting expression, training a human to go through millions of protein sequences to find relevant patterns is not feasible, but can easily be automated by AI (Avsec et al., 2021).
The advent of these tools will improve the availability of protein stably expressed and purified with high yields, which is crucial to the success of structural biology.
4. Automating Data Collection for X-ray Crystallography and Cryo-EM
AI tools are being developed to deal with big data generated, for example by serial crystallography data collection at conventional synchrotron, or Free Electron Laser sources (Vollmar and Evans, 2021). Other AI approaches enable to cope with very large dataset and make Cryo-EM more feasible with efficient use of computational resources. For instance, AI-empowered reconstruction of 3D volume from 2D images using prior knowledge can provide better classification, higher resolution and accelerate data analysis (Zhong et al., 2021).
AI tools can assist experimentalists and improve the resolution of protein structures, which in turn, is providing better information to drive the design of compounds.
5. Predicting Polypharmacology and Finding New Targets
AI is also changing the way targets are selected and validated. It becomes possible to find promising targets by integrating data from different sources: patient populations omics data, silencing RNA, animal models, clinical trials, patents, and literature data. AI can also estimate the likelihood of a molecule to hit other targets – by learning binding patterns from the aligned binding sites of all solved protein structure complexes – which can be used for drug repositioning or for better anticipating side effects and toxicity (Chaudhari et al., 2020).
Here, the availability of a comprehensive map of all human proteins of therapeutic relevance together with 3-dimensional structural information can greatly facilitate the drug discovery process.
In summary, the interface between structural biology and AI will continue to be fertile ground for breakthrough technologies that are already today accelerating the discovery of future medicines.
References
Avsec, Ž., Agarwal, V., Visentin, D., Ledsam, J.R., Grabska-Barwinska, A., Taylor, K.R., Assael, Y., Jumper, J., Kohli, P., Kelley, D.R., 2021. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203. https://doi.org/10.1038/s41592-021-01252-x
Botte, M., Ni, D., Schenck, S. et al. Cryo-EM structures of a LptDE transporter in complex with Pro-macrobodies offer insight into lipopolysaccharide translocation. Nat Commun 13, 1826 (2022). https://doi.org/10.1038/s41467-022-29459-2 – see also our news article
Chan, H.C.S., Shan, H., Dahoun, T., Vogel, H., Yuan, S., 2019. Advancing Drug Discovery via Artificial Intelligence. Trends Pharmacol. Sci., Special Issue: Rise of Machines in Medicine 40, 592–604. https://doi.org/10.1016/j.tips.2019.06.004
Chaudhari, R., Fong, L.W., Tan, Z., Huang, B., Zhang, S., 2020. An up-to-date overview of computational polypharmacology in modern drug discovery. Expert Opin. Drug Discov. 15, 1025–1044. https://doi.org/10.1080/17460441.2020.1767063
de Almeida, A.F., Moreira, R., Rodrigues, T., 2019. Synthetic organic chemistry driven by artificial intelligence. Nat. Rev. Chem. 3, 589–604. https://doi.org/10.1038/s41570-019-0124-0
Hopf, T.A., Green, A.G., Schubert, B., Mersmann, S., Schärfe, C.P.I., Ingraham, J.B., Toth-Petroczy, A., Brock, K., Riesselman, A.J., Palmedo, P., Kang, C., Sheridan, R., Draizen, E.J., Dallago, C., Sander, C., Marks, D.S., 2019. The EVcouplings Python framework for coevolutionary sequence analysis. Bioinforma. Oxf. Engl. 35, 1582–1584. https://doi.org/10.1093/bioinformatics/bty862
Jumper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., Tunyasuvunakool, K., Bates, R., Žídek, A., Potapenko, A., Bridgland, A., Meyer, C., Kohl, S.A.A., Ballard, A.J., Cowie, A., Romera-Paredes, B., Nikolov, S., Jain, R., Adler, J., Back, T., Petersen, S., Reiman, D., Clancy, E., Zielinski, M., Steinegger, M., Pacholska, M., Berghammer, T., Bodenstein, S., Silver, D., Vinyals, O., Senior, A.W., Kavukcuoglu, K., Kohli, P., Hassabis, D., 2021. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589. https://doi.org/10.1038/s41586-021-03819-2
Olivecrona, M., Blaschke, T., Engkvist, O., Chen, H., 2017. Molecular de-novo design through deep reinforcement learning. J. Cheminformatics 9, 48. https://doi.org/10.1186/s13321-017-0235-x
Paul, D., Sanap, G., Shenoy, S., Kalyane, D., Kalia, K., Tekade, R.K., 2021. Artificial intelligence in drug discovery and development. Drug Discov. Today 26, 80–93. https://doi.org/10.1016/j.drudis.2020.10.010
Popov, P., Peng, Y., Shen, L., Stevens, R.C., Cherezov, V., Liu, Z.-J., Katritch, V., 2018. Computational design of thermostabilizing point mutations for G protein-coupled receptors. eLife 7, e34729. https://doi.org/10.7554/eLife.34729
Schneider, G., Fechner, U., 2005. Computer-based de novo design of drug-like molecules. Nat. Rev. Drug Discov. 4, 649–663. https://doi.org/10.1038/nrd1799
Struble, T.J., Alvarez, J.C., Brown, S.P., Chytil, M., Cisar, J., DesJarlais, R.L., Engkvist, O., Frank, S.A., Greve, D.R., Griffin, D.J., Hou, X., Johannes, J.W., Kreatsoulas, C., Lahue, B., Mathea, M., Mogk, G., Nicolaou, C.A., Palmer, A.D., Price, D.J., Robinson, R.I., Salentin, S., Xing, L., Jaakkola, T., Green, William.H., Barzilay, R., Coley, C.W., Jensen, K.F., 2020. Current and Future Roles of Artificial Intelligence in Medicinal Chemistry Synthesis. J. Med. Chem. 63, 8667–8682. https://doi.org/10.1021/acs.jmedchem.9b02120
Vollmar, M., Evans, G., 2021. Machine learning applications in macromolecular X-ray crystallography. Crystallogr. Rev. 27, 54–101. https://doi.org/10.1080/0889311X.2021.1982914
Zhong, E.D., Lerer, A., Davis, J.H., Berger, B., 2021. CryoDRGN2: Ab Initio Neural Reconstruction of 3D Protein Structures From Real Cryo-EM Images. Presented at the Proceedings of the IEEE/CVF International Conference on Computer Vision, pp. 4066–4075. https://openaccess.thecvf.com/content/ICCV2021/papers/Zhong_CryoDRGN2_Ab_Initio_Neural_Reconstruction_of_3D_Protein_Structures_From_ICCV_2021_paper.pdf