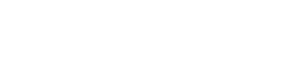First structure of human Kv3.1 channel
In collaboration with Lundbeck, leadXpro solved the first structure of a human Kv3 channel in complex with a positive modulator.
Apo and ligand-bound high resolution Cryo-EM structures of the human Kv3.1 reveal a novel binding-site for positive modulators.
leadXpro has solved the Kv3.1 by cryo-EM, at 2.6 Angstrom, in the apo state and in a complex with a positive modulator designed by Lundbeck (Lu AG00563). This represents the first structure of a Kv3 channel, of which many members are implicated in diseases, in particular in the CNS space.
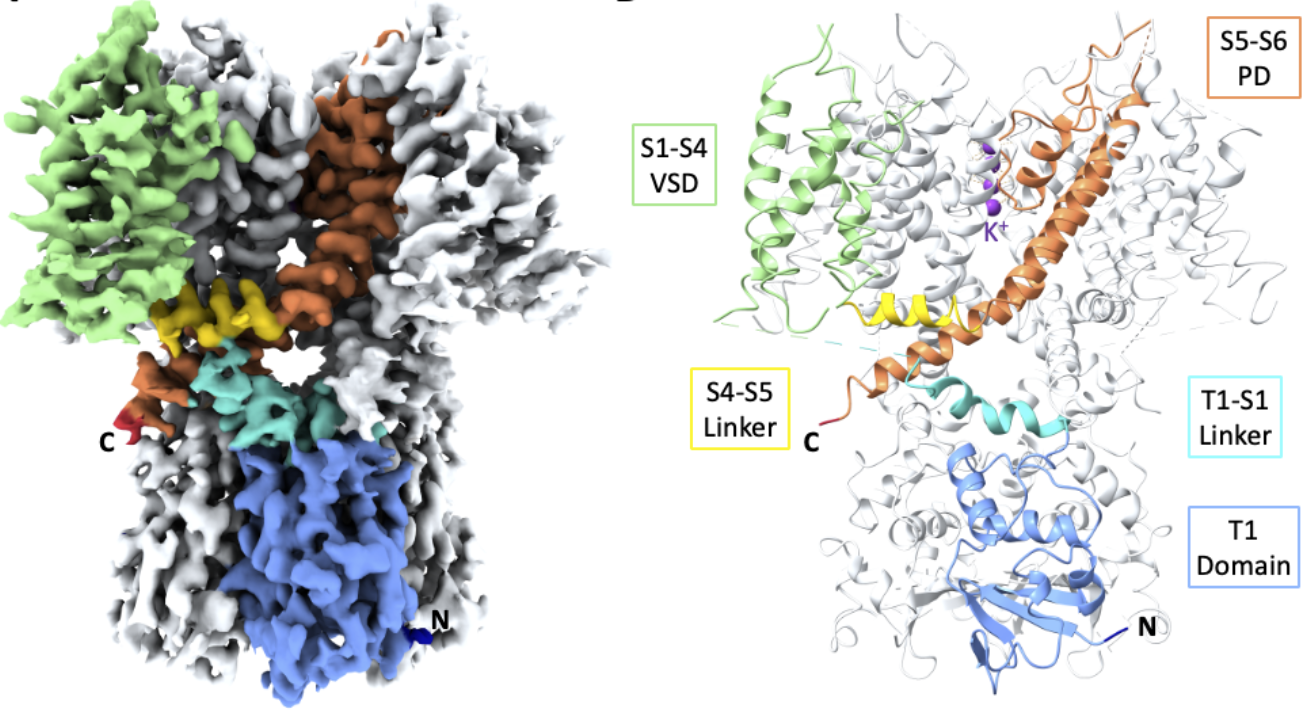
Of particular interest is the discovery of a zinc binding site that has been hypothesized to participate in the channel regulation. The zinc atom is seen to stabilize a rotated conformation by 35 deg of the cytoplasmic T1 domain which is not seen in other Kv classes.
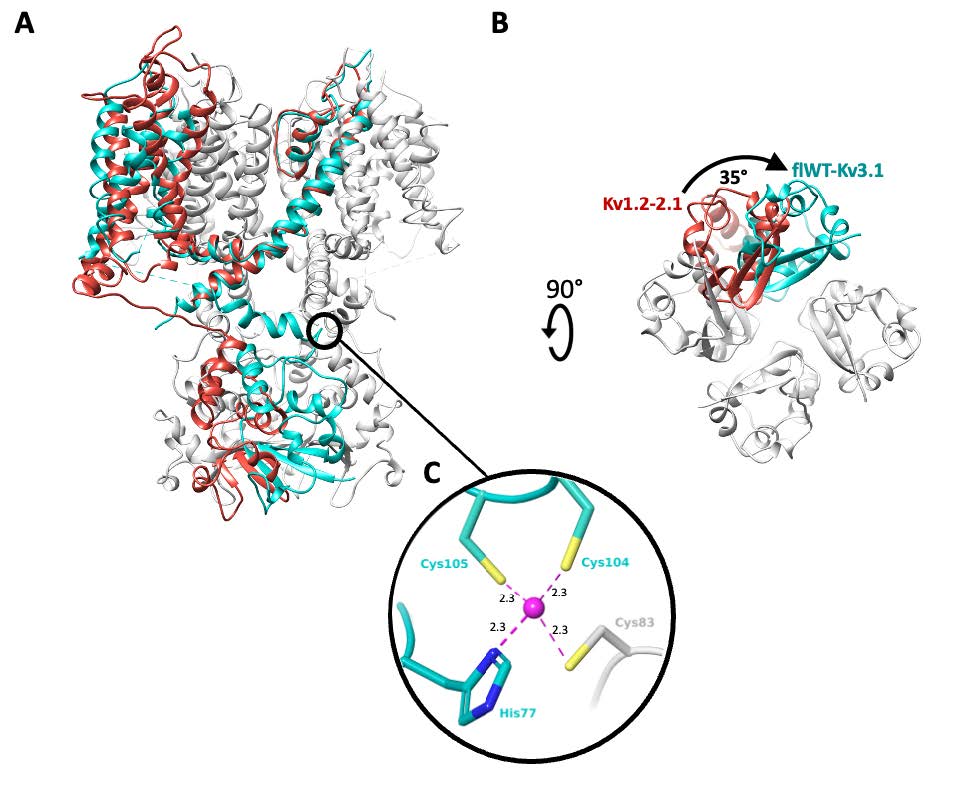
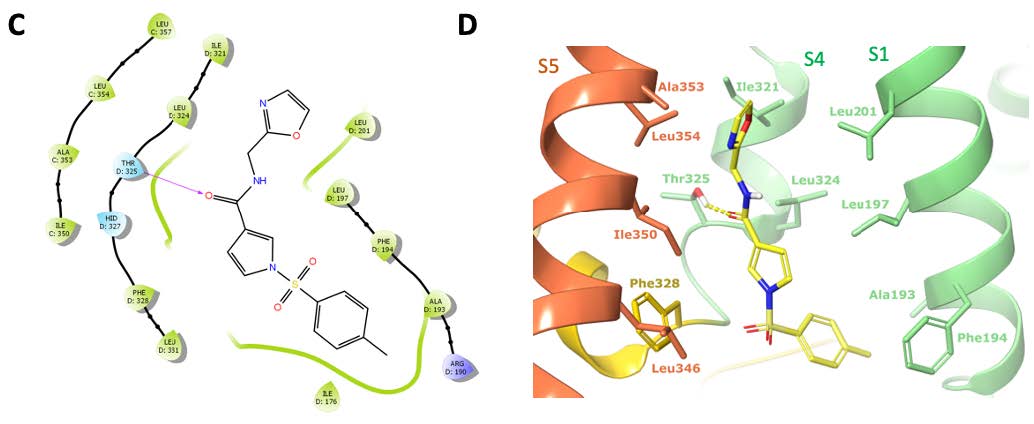
In addition, the binding site identified for Lu AG00563 was found to be unique in all related Kv structures and located between the voltage sensory domain and the channel pore, in a region known to be a hotspot for disease-causing mutations. The knowledge of such a site for positive modulators could now set the stage for future structure-based drug discovery programs.
Reference:
Mathieu Botte, Sophie Huber, Denis Bucher, Julie K Klint, David Rodríguez, Lena Tagmose, Mohamed Chami, Robert Cheng, Michael Hennig, Wassim Abdul Rahman: Apo and ligand-bound high resolution Cryo-EM structures of the human Kv3.1 channel reveal a novel binding site for positive modulators, PNAS Nexus, Volume 1, Issue 3, July 2022, pgac083, https://doi.org/10.1093/pnasnexus/pgac083